More than 64,000 proteomics datasets person now flowed done ProteomeXchange, and nan consortium’s latest update shows really smarter standards, stronger reuse tools, and AI-ready resources are reshaping biologic information sharing.
 Database Update: The ProteomeXchange consortium successful 2026: making proteomics information FAIR. Image Credit: Christoph Burgstedt /Shutterstock
Database Update: The ProteomeXchange consortium successful 2026: making proteomics information FAIR. Image Credit: Christoph Burgstedt /Shutterstock
In a caller database update insubstantial published successful nan journal Nucleic Acids Research, an world squad of authors described caller advancements, information growth, standardization, and early directions of nan ProteomeXchange Consortium successful enabling FAIR (Findable, Accessible, Interoperable, Reusable) proteomics information sharing.
Proteomics Data Sharing Background and FAIR Principles
What happens erstwhile thousands of biologic datasets stay unused? In proteomics, information sharing is basal to beforehand investigation connected diseases, drugs, and quality biology. Over nan past decade, nan accelerated emergence of wide spectrometry-based proteomics has generated immense datasets, yet their worth depends connected accessibility and reuse. The FAIR principles were developed to guideline technological information guidance and stewardship successful ways that support reproducible and transparent science. Collaborative platforms now play a important domiciled successful integrating and distributing specified information crossed disciplines. However, continuous invention is needed to grip nan increasing complexity of caller datasets.
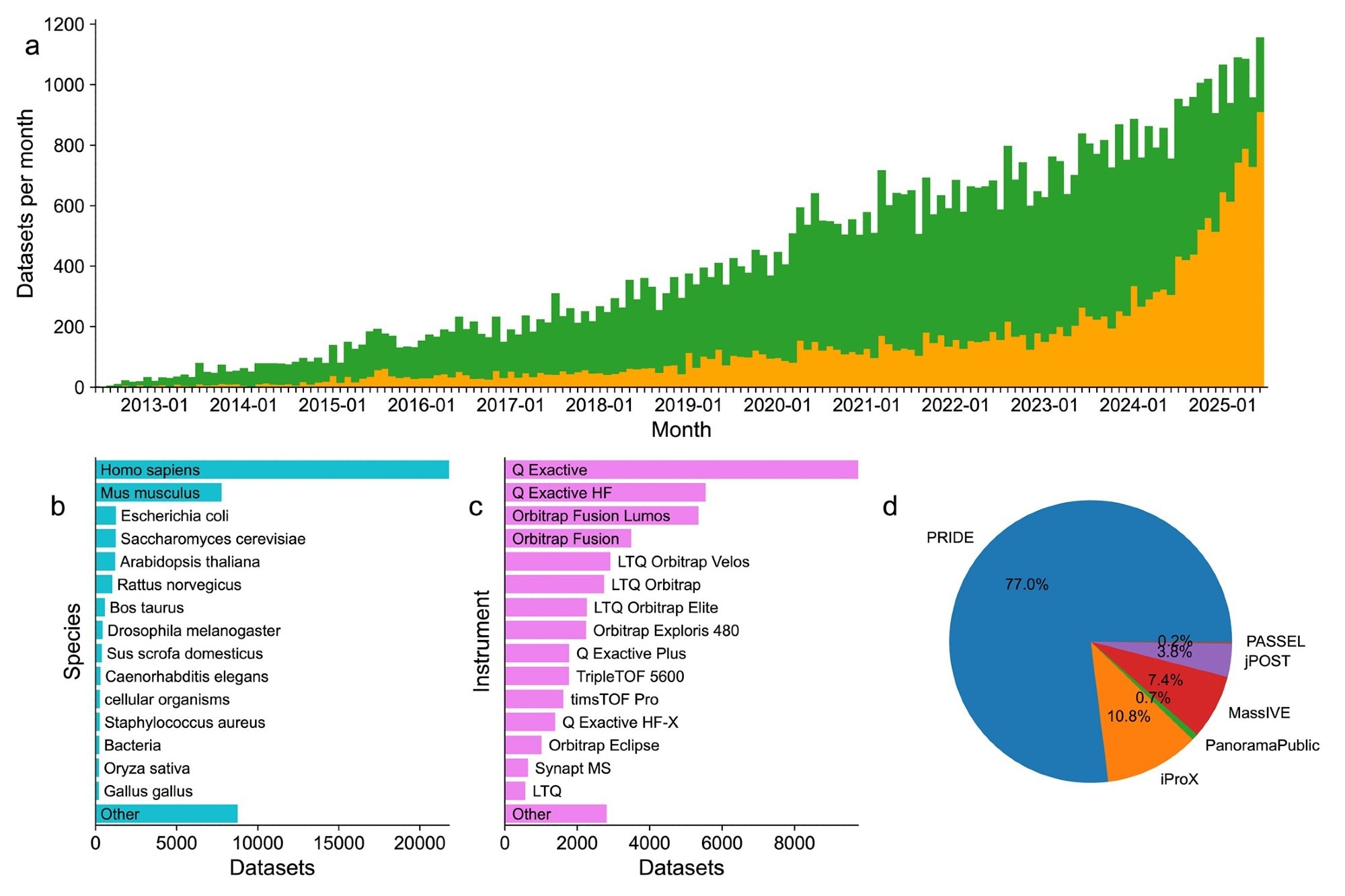
Summary statistic for datasets deposited to ProteomeXchange resources since 2012. (A) Trend successful publically released (green) and not-yet released (orange) datasets from May 2012 done June 2025. A full of 1156 datasets were submitted successful June 2025. (B) Summary of nan apical 15 type for publically released datasets since 2012. (C) Summary of nan apical 15 instruments arsenic reported by submitters for publically released datasets since 2012. (D) Summary of nan comparative number of each datasets by nan receiving repository.
ProteomeXchange Infrastructure and Data Standards
The consortium maintains an infrastructure that allows for nan standardized submission, storage, and dissemination of proteomic information generated by wide spectrometry. Member repositories that contributed to information archiving and entree see PRoteomics IDEntifications database (PRIDE), PeptideAtlas, Mass Spectrometry Interactive Virtual Environment (MassIVE), Japan Proteome Standard Repository/Database (jPOST), Integrated Proteome Resources (iProX), and Panorama Public. Datasets submitted consisted of earthy wide spectrometry files, processed information pinch recognition and quantification results, and experimental metadata system according to Proteomics Standards Initiative (PSI)-developed standards.
Efficient uploads were conducted utilizing a number of information transportation protocols, including File Transfer Protocol (FTP), Aspera, Hypertext Transfer Protocol Secure (HTTPS), Web Distributed Authoring and Versioning (WebDAV), and PRESTO. Additionally, standardization of metadata was improved done nan Sample and Data Relationship Format (SDRF)-Proteomics, enabling clear mapping betwixt samples and experimental conditions. Unique dataset identifiers (ProteomeXchange dataset identifiers) ensured traceability, while reanalyzed datasets were assigned RPXD identifiers.
ProteomeCentral integrated metadata from each repositories, enabling hunt and retrieval of datasets done a azygous platform. Universal Spectrum Identifiers (USIs) allowed for meticulous recognition and visualization of azygous spectra. The infrastructure besides facilitated their reuse astatine scale, integration pinch outer resources, and usage successful instrumentality learning and artificial intelligence (AI) workflows.
ProteomeXchange Growth, Reuse, and AI Applications
Updated submission statistic from nan consortium showed important maturation successful world proteomics information sharing and reuse. By June 2025, a full of 64,330 datasets had been submitted, pinch 44,248 (69%) publically accessible, reflecting a beardown committedness to unfastened science. Notably, 47% of each datasets were submitted wrong nan past 3 years, highlighting an accelerating inclination successful information procreation and sharing.
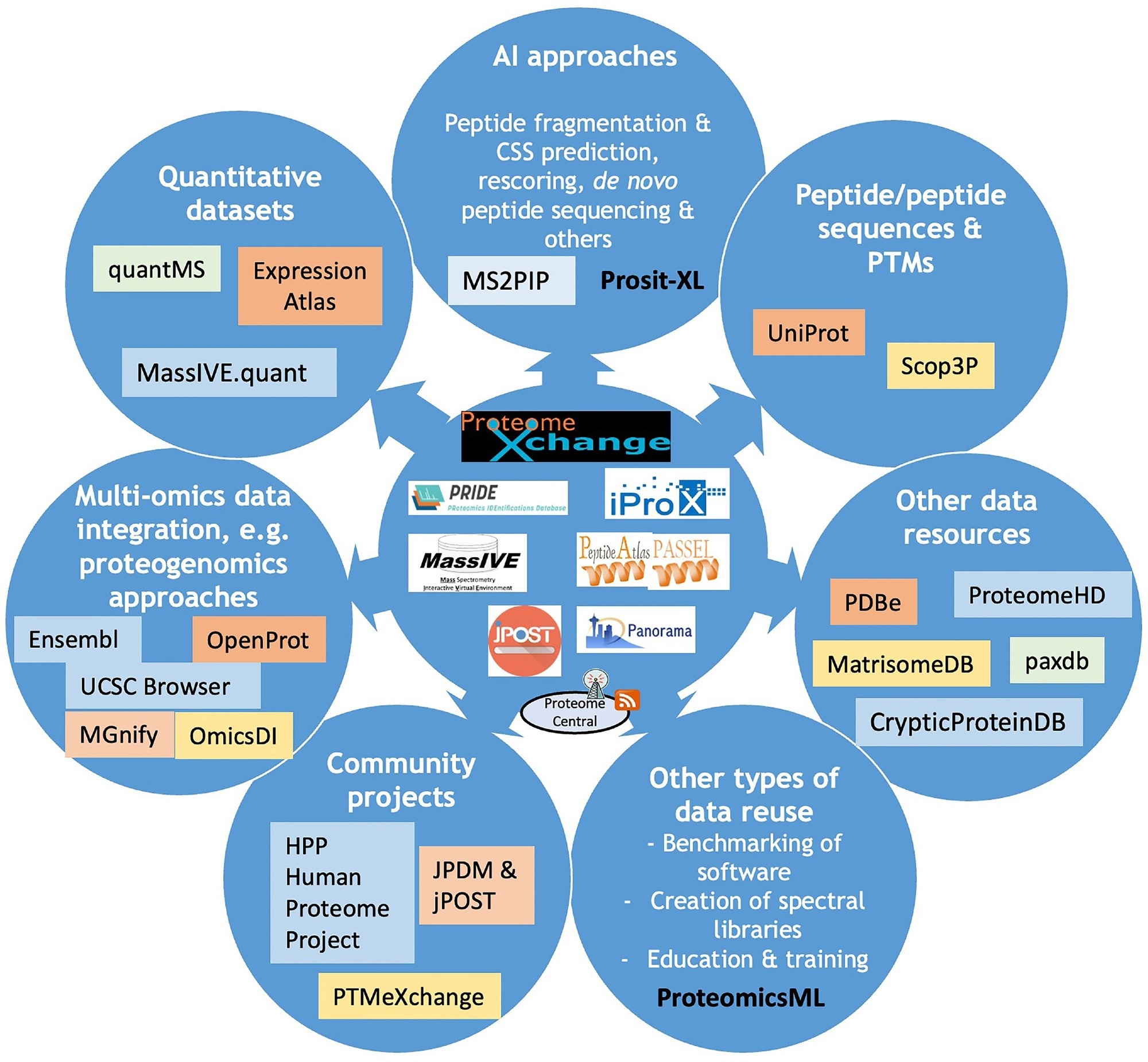
Overview fig including nan existent ProteomeXchange resources and nan main efforts devoted to information reuse of nationalist proteomics datasets. Different types of information reuse are listed and for each of them, nan corresponding devices and/or information resources wherever these information tin beryllium accessed are indicated.
Most of nan submissions were from nan PRIDE repository (77%), followed by iProX (11%), MassIVE (7.4%), jPOST (3.8%), and very mini amounts from Panorama Public and PeptideAtlas. Over 80 countries contributed to these nationalist proteomics resources, indicating that proteomics usage successful biomedical investigation is wide globally.
ProteomeXchange resources progressively support standardized formats and richer metadata to heighten interoperability crossed datasets. The PSI-developed formats and SDRF-Proteomics enhanced nan metadata from nan datasets by improving their quality, reproducibility, and value. The wide usage of USIs facilitated nan entree to and visualization of individual spectra successful aggregate different information repositories. This enhanced nan transparency and validation of experimental results.
Data reuse activities besides accrued crossed nan consortium. Public datasets were reanalyzed to get caller biologic insights, specified arsenic validating macromolecule sequences and identifying post-translational modifications. The integration pinch UniProt Knowledge Base (UniProtKB) helped representation much than 93% of nan quality proteome, showing nan powerfulness of information analytics.
Quantitative proteomics resources specified arsenic MassIVE.quant and quantms enabled reproducible large-scale analyses. Additionally, multi-omics integration done resources for illustration Omics Discovery Index (OmicsDI) and MGnify helped merge proteomics, genomics, and transcriptomics datasets.
Artificial intelligence and instrumentality learning applications were progressively supported by nan readiness of high-quality datasets. Tools specified arsenic MassIVE-Knowledge-Base (MassIVE-KB) and ProteomicsML enabled nan improvement of predictive models for peptide identification, fragmentation, and macromolecule quantification. These advances are transforming proteomics into a data-driven section pinch imaginable early applications successful precision medicine.
There are still galore challenges that beryllium successful this section of research. Due to privateness regulations for illustration nan General Data Protection Regulation (GDPR) and nan Health Insurance Portability and Accountability Act (HIPAA), much controlled-access systems and repository capabilities are needed for quality data. Additionally, caller technologies person emerged that usage proteomics arsenic a superior measurement method and do not dangle connected wide spectrometry, including affinity proteomics platforms specified arsenic SomaLogic and Olink assays. This will lead to caller investigation methodologies; therefore, researchers whitethorn request further resources.
Future Directions for FAIR Proteomics Infrastructure
The ProteomeXchange Consortium has created an innovative, collaborative situation for nan world sharing of proteomics data, aligned pinch FAIR principles. The preamble of standardized formats, accrued scalability, and nan proviso of cutting-edge analytical devices person facilitated nan wide reuse of existing information to beforehand innovations successful biology and medicine. However, early advancement depends connected solving information privacy, scalability, and emerging technologies.
There is an ongoing request for invention and collaboration to support wide accessibility and support nan continued reliability and effect of proteomics information successful advancing technological find and enabling wider bioinformatics reuse.
Source:
Journal reference:
- Deutsch, E. W., Bandeira, N., Perez-Riverol, Y., Sharma, V., Carver, J. J., Mendoza, L., Kundu, D. J., Bandla, C., Kamatchinathan, S., Hewapathirana, S., Sun, Z., Kawano, S., Okuda, S., Connolly, B., MacLean, B., MacCoss, M. J., Chen, T., Zhu, Y., Ishihama, Y., & Vizcaíno, J. A. (2026). The ProteomeXchange consortium successful 2026: Making proteomics information FAIR. Nucleic Acids Research. 54(D1). D459–D469. DOI: 10.1093/nar/gkaf1146, https://academic.oup.com/nar/article/54/D1/D459/8315797
.png?2.1.1)







 English (US) ·
English (US) ·  Indonesian (ID) ·
Indonesian (ID) ·